A Princeton University-led research team has discovered that protein competition over an important enzyme provides a mechanism to integrate different signals that direct early embryonic development. The work suggests that these signals are combined long before they interact with the organism's DNA, as was previously believed, and also may inform new therapeutic strategies to fight cancer.
The fought-over enzyme, known as the mitogen-activated protein kinase (MAPK), is found in all complex organisms, ranging from yeast to humans. MAPK signaling pathways, or chemical networks that involve the enzyme, are critical for normal development, and defects in these pathways can lead to severe developmental disorders and cancer.
During early embryonic development, a single undifferentiated cell becomes a complex and highly specialized organism containing a variety of different cell types arranged in very precise patterns. These patterns, which ensure that the body structures from head-to-tail and front-to-back develop correctly and in the appropriate places, are created when cells respond to a series of chemical signals from different signaling pathways. The different patterning signals received by any given cell are ultimately combined to govern its future fate and tell it what kind of cell it should become.
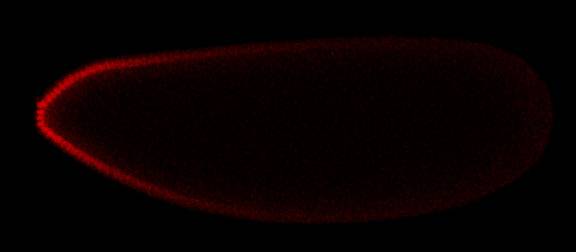
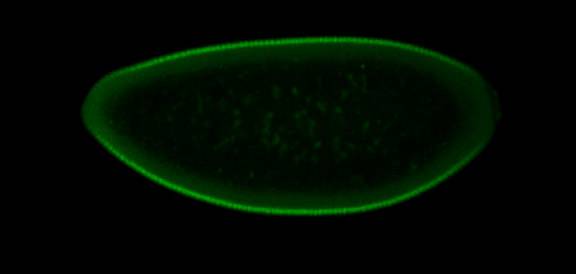
The Princeton team used confocal microscopy to visualize the spatial distribution of two proteins that compete for the MAPK enzyme in early fruit fly embryos. In areas where levels of a protein important for the development of the head were high (shown here in red, with brighter color indicating the presence of more protein) there was less enzyme available to act upon a different protein (shown here in green) that is important for the development of the ends of the embryo, including the tail. (Images: Shvartsman Lab)Images for news media
Until now, scientists believed these pathways operated largely independently of one another to produce protein signals that traveled to the nuclei of the embryo's cells where DNA is stored. There, coordination of these signals was thought to occur when they interacted with cell DNA to influence and control the expression of genes. Results published March 9 in the journal Current Biology, however, suggest that competition for the MAPK enzyme among proteins in different pathways influences which signals are sent to cells, establishing a biochemical mode of signal integration that adds a previously unrecognized layer of complexity and control to embryonic development.
"It appears that different proteins in different pathways are competing for the MAPK enzyme inside these living organisms," said Stanislav Shvartsman, associate professor in the Department of Chemical Engineering and the Lewis-Sigler Institute for Integrative Genomics who earned his Ph.D. from Princeton in 1999. "Since these proteins are fighting for the same limited resource -- the enzyme -- they indirectly control one another, which in turn coordinates the developmental signals."
Conventional biology teaches that enzymes like MAPK act on certain molecules, called substrates, to regulate chemical reactions. The new findings are surprising because it appears that, through competition with one another, the substrates of MAPK are, in fact, influencing the enzyme's activity.
"In a way, it's like the tail wagging the dog," Shvartsman said. "The substrates are regulating the enzyme, and, by extension, mediating the chemical reactions."
Eric Wieschaus, Princeton’s Squibb Professor in Molecular Biology who received the 1995 Nobel Prize in medicine for his pioneering work in developmental biology, said, "Their results argue convincingly that these signaling molecules are interacting with each other in a competitive way such that even before anything gets to the DNA, they’ve already made decisions. Essentially the decisions aren’t just made in terms of DNA, but also in terms of proteins working together. This is, in a way, revolutionary."
The research team, led by Princeton chemical engineering graduate student Yoosik Kim, focused its attention on the interaction between MAPK and two proteins involved in two different signaling pathways for head-to-tail pattern formation. The first of these proteins is part of the pathway that governs the development of the head. The second protein plays a significant role in the chemical circuit that controls the development of the ends of the embryo, including the tail.
Using special techniques to visualize whether the proteins had interacted with the MAPK enzyme, the team found that the relative amount of the first protein controlled how much enzyme was available to interact with the second protein. For example, in the portion of the embryo that would become the head, where the concentration of the first protein was high, much less enzyme was available to act on the second protein than at the other end of the embryo, where the tail would ultimately develop.
"This competition makes sure that the same enzyme signals are interpreted differently in the head and in the tail, thereby allowing for the integration of multiple signals," Shvartsman said.
Based on how the enzyme interacted with the proteins in the head region of the embryo, the team predicted that a third protein also might be competing for the MAPK enzyme in that area. To test the hypothesis, research team members at the Institute for Medical Research Israel-Canada at Hebrew University in Jerusalem used a series of experimental techniques to verify that their proposed protein could bind to the enzyme, an ability that was previously unknown. These findings suggest that the competition model may provide a novel way to identify proteins that are involved in signaling pathways.
Beyond advancing the fundamental understanding of mechanisms that control embryonic patterning, the work has implications for how to target cancer cells, which often exhibit hyperactive MAPK signaling.
"According to our substrate competition idea, MAPK signaling activity directed toward any given substrate decreases when you introduce a competing substrate," Kim said. "In theory, you can lower the activity of MAPK if you introduce a protein whose sole function is to bind to MAPK and thus act as a competitive inhibitor of MAPK signaling to all other substrates." This strategy might one day allow scientists to slow or stop MAPK signaling pathways in cancer cells by adding a protein that monopolizes the MAPK enzyme, effectively disrupting the chemical circuitry of a cancer cell.
In future work, the researchers plan to conduct experiments to investigate competition among other proteins that bind to MAPK and to investigate how this competition for the MAPK enzyme manifests itself in other organisms. The group also intends to explore how certain proteins are able to outcompete other proteins for the enzyme's attention, perhaps by binding more strongly or efficiently to the molecule. In time, the group may expand its work to consider whether similar competition models affect the activity of different enzymes in other signaling pathways.
In addition to Kim and Shvartsman, the team included Princeton research associate Matthieu Coppey; Rona Grossman and Ze’ev Paroush of the Institute for Medical Research Israel-Canada; and Leiore Ajuria and Gerardo Jimenez of the Barcelona Institute for Molecular Biology. The research was supported by the U.S. National Institutes of Health, the Israel Science Foundation, the Israel Cancer Research Fund, the Krol Charitable Foundation, the Spanish Ministry of Science and Education, and the Institucio Catalana de Recerca i Estudis Avancats.