New research suggests that just one or two individual herpes virus particles attack a skin cell in the first stage of an outbreak, resulting in a bottleneck in which the infection may be vulnerable to medical treatment.
Unlike most viruses that spread to new cells by bombarding them with millions of particles, herpes simplex virus type 1 (HSV-1) — a virus that causes cold sores and genital lesions — requires just one or two viral particles to infect a skin cell in the first stage of cold sore formation, Princeton University researchers reported in the Proceedings of the National Academy of Sciences.
"The fact that just one or two virus particles are involved is surprising because these viruses can replicate themselves hundreds of times in a single cell," said Matthew Taylor, first author on the study and a postdoctoral researcher in the laboratory of Lynn Enquist, the Henry L. Hillman Professor in Molecular Biology and the Princeton Neuroscience Institute.
The bottleneck occurs when HSV-1 particles, which can lie dormant in the cells of the nervous system for decades after initial infection, awaken and invade a nearby skin cell, the first stage in sore formation. Once inside the skin cell, a single viral particle multiplies and spreads millions of copies to nearby skin cells, creating a visible lesion or "cold sore." The virus can then spread to new individuals through skin-to-skin contact.
This restriction to one or two particles limits the genetic diversity of the virus that spreads to the next individual, Enquist said. This puts the virus at a disadvantage, he said, because having a variety of distinct genomes enhances the overall chances of the virus surviving and spreading. A well-known example of a virus that relies on genetic diversity to thrive is HIV, which involves large numbers of viral particles with distinct genomes.
In the case of herpes viruses, Enquist said: "The number of different genomes that infect the cell is remarkably low, so any mutations that weaken the virus are unlikely to survive. Only the most fit viral particles will survive and replicate in the epithelial [skin] cells, and be available to transmit to the next individual."
Although bottlenecks can ensure that only the most fit viruses are transmitted to the next individual, they also can be points at which the infection is more susceptible to immune system responses and drug treatments, according to Taylor. He and Enquist worked with co-author Oren Kobiler, a former Princeton postdoctoral researcher now at Tel Aviv University.
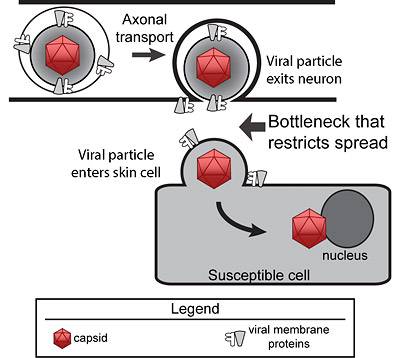
Model of herpes virus spread from neuron to skin cell Herpes virus particles move via axonal transport inside a neuron. Upon exiting the neuron, only one or two viral particles enter the susceptible skin cell, resulting in a bottleneck that restricts viral spread. (Image courtesy of Matthew Taylor)
The researchers' findings suggest that other viruses related to HSV-1 — known as alpha-herpes viruses — may have similar bottlenecks, Taylor said, including herpes simplex viruses type 2, which causes cold sores and genital lesions, and varicella zoster virus, which causes chicken pox and shingles. It remains to be determined if this bottleneck exists for other viruses that spread from infected neurons, such as poliovirus and the West Nile virus, Taylor said.
Julie Pfeiffer, an associate professor of microbiology at the University of Texas Southwestern Medical Center, said the Princeton research is the first to tally how many viral particles are involved in HSV-1 infection — and reveal that as the virus' weak point.
"This work changes the way that we think about herpes virus spread," said Pfeiffer, who is familiar with the study but had no role in it. "This study demonstrated that these neuron-to-epithelial cell viral-transmission events are surprisingly efficient, but they are initiated by a very small number of viruses. This work has interesting implications for herpesvirus transmission and evolution."
To determine the number of virus particles that infect the skin cell, the researchers constructed three genetically unique viral genomes labeled with either red, green or blue fluorescent tags, and then infected cells with the particles and analyzed them for the presence of the three colors. They counted the number of cells containing one, two or all three colors and used statistical analysis to determine the number of unique viral genomes expressed in each cell. The method, which was originally developed by Kobiler, Enquist and colleagues in Princeton's mathematics department and initially published in 2010 in Nature Communications, revealed that most skin cells express less than two viral genomes on average.
Taylor then filmed individual virus particles of pseudorabies virus, a model alpha-herpes virus that infects animals, as the particles exited neurons and entered skin cells. He found that the bottleneck limiting infection to one or two particles exists for pseudorabies virus as well.
As shown in the video of the process (above), viral particles, genetically altered to glow red, travel down long extensions of the neuron cell body called axons.
The video, courtesy of Taylor, reveals a handful of viral particles exiting an axon but only one particle entering a nearby epithelial cell. Once inside, the virus particle replicates and creates millions of particles that fill up the cell, turning it red in color. These viral particles later exit the infected cell and attack surrounding epithelial cells.
"It is really amazing to see this happening in living cells," Taylor said.
The paper, "Alphaherpesvirus axon-to-cell spread involves limited virion transmission," was published Oct. 16 in the Proceedings of the National Academy of Sciences. The work was supported by the National Institutes of Health and the American Cancer Society.