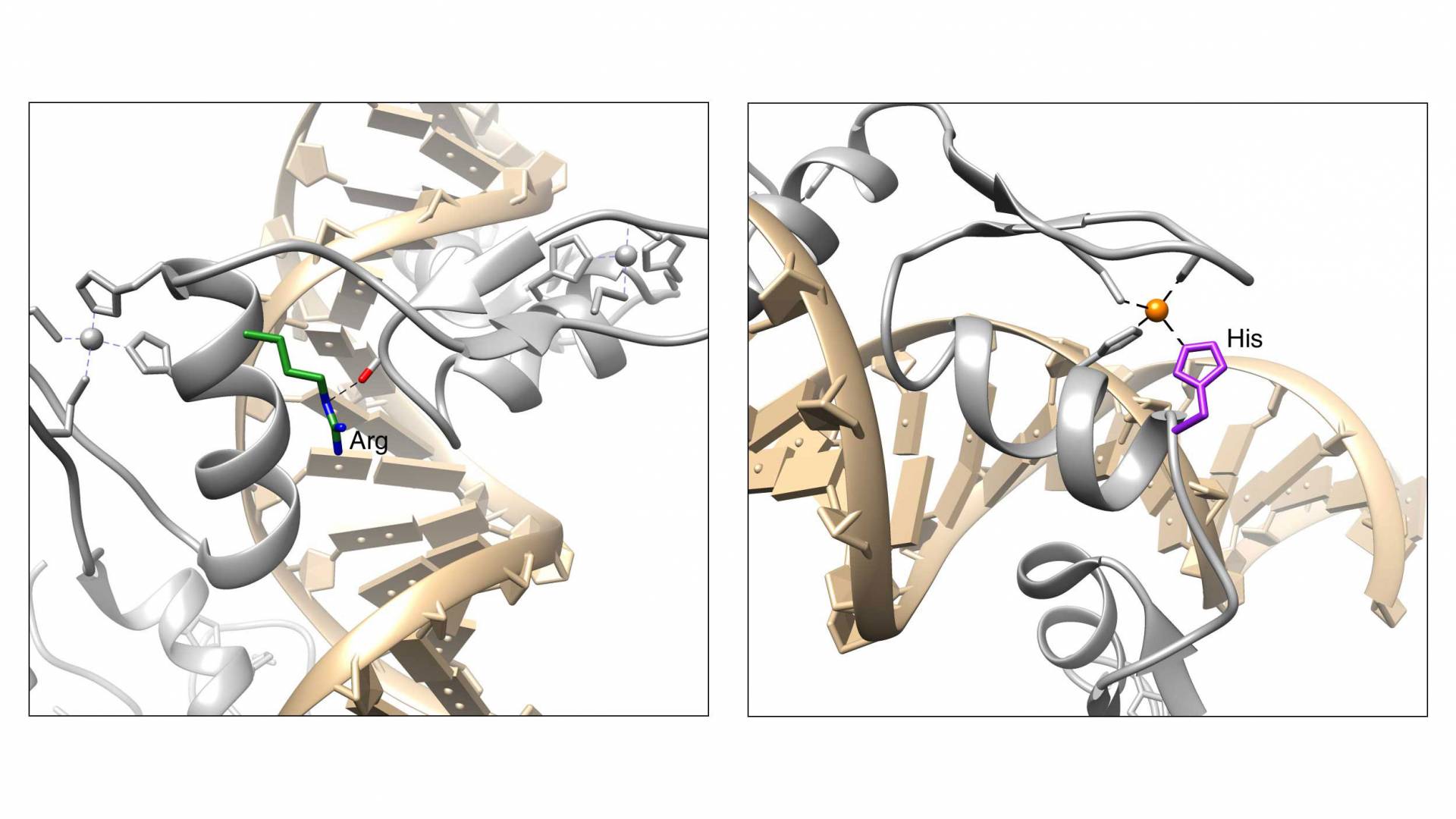
By analyzing data from thousands of patients, Princeton researchers led by Professor Mona Singh have identified two genetic mutations occurring at high frequencies in three types of cancer. The mutations affect amino acids in protein molecules shown in the panels above.
By analyzing data from thousands of patients, Princeton researchers have identified genetic mutations that frequently occur in people with uterine cancer, colorectal cancer or skin cancer — an important step toward using genome sequences to better understand cancer and guide new treatments.
Many genetic malfunctions are commonly found in cancers, and scientists are working to distinguish those directly involved in both the initial growth of cancer and its spread throughout the body. The complexity of both the disease and the ordinary functions of the genome have made this a difficult problem. However, researchers have been applying advances in data science to new collections of genetic data to the question of how cancer develops.
“There are numerous mutations that occur in any individual’s cancer,” said Mona Singh, senior author of the paper and a Princeton professor of computer science and the Lewis-Sigler Institute for Integrative Genomics. “A major question in cancer genomics is to figure out, from this large sea of mutations, which ones are actually causal for cancer initiation, but also for progression.”
In a study published June 28 in the journal PLOS Computational Biology, Singh’s group addressed this question by lining up mutations in a large group of genes that play key roles in gene expression regulation and other cellular processes. These genes are known as zinc finger genes because they have a protein structure that involves a zinc ion. The researchers examined mutations in these genes using the Cancer Genome Atlas, which includes more than 10,000 tumor samples from 32 cancer types.
Daniel Munro, the paper’s lead author, said the researchers were able to take advantage of the genes’ similarities to find common cancer mutations that disrupt protein structures, leading to impaired cellular functions.
“It allows us to identify patterns because we can look at many different proteins at the same time,” said Munro, a graduate student in the Lewis-Sigler Institute for Integrative Genomics. Dario Ghersi, a former postdoctoral researcher in Singh’s lab who is now an assistant professor at the University of Nebraska Omaha, was also a co-author.
Cancer cells divide rapidly and often have deficient DNA repair mechanisms, leading to high mutation rates compared to normal cells. Therefore the researchers tested whether the mutations they had detected occurred more often than would be expected by chance — an important check to evaluate the meaning of their results. They calculated the overall mutation rates for zinc finger genes in each cancer patient and found that two sites — affecting amino acids 9 and 11 in the zinc finger domain — were mutated at higher rates than this background level.
Among other control analyses, the team also compared the mutation rates at amino acids 9 and 11 to rates of “mutational signatures” known to be common across the genome in particular cancer types. Although such mutations are common in these cancers, at amino acid 9 in zinc finger domains they occurred nearly 10 times more often than would be expected by chance in uterine and colorectal cancers.
The study further showed that in other cancer types, zinc finger genes have more mutations than the genome overall. Taken together, the results are strong evidence “that zinc finger genes are major players in the gene dysregulation you see in cancers,” said Singh.
Singh said she hoped the results would help guide laboratory research and lead to progress in understanding the mechanisms of cancer.
“We’re keen to have other people follow up on this work,” she said.
The research was supported in part by the National Science Foundation, the National Institutes of Health, the State of Nebraska and the Nebraska Research Initiative.